The Genetic Piece Of The GLP-1 Response Puzzle
Inside the largest genetic study of GLP-1 response to date
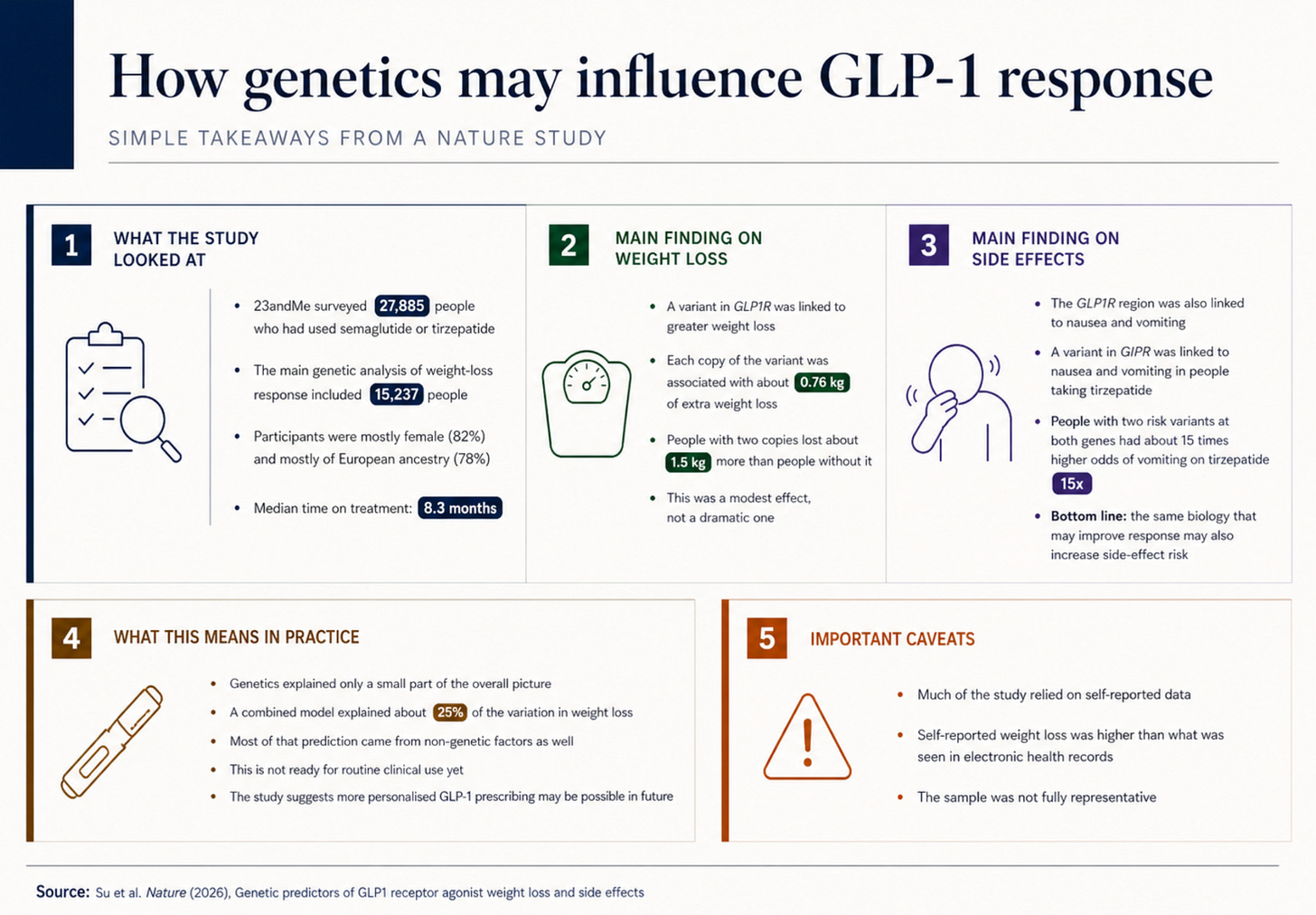
A new study published in Nature has identified two gene variants that influence how people respond to GLP-1 medications. It’s one of the largest pharmacogenomic studies in this space to date and the findings are genuinely interesting.
The problem we already know
GLP-1 response varies enormously. Across studies, a meaningful minority of people do not reach the typical response threshold of at least 5% body weight loss. In one semaglutide analysis, 32% of people achieved less than 5% loss from baseline or gained weight, while 5% lost more than 25%. That’s a wide disparity in outcomes from the same class of drug.
What this study found
Researchers at the 23andMe Research Institute ran a genome wide association study across 27,885 people who had used semaglutide or tirzepatide.
Two key findings emerged:
A variant in the GLP1R gene (the gene encoding the receptor these drugs target) was associated with greater weight loss. Each copy of the variant contributed roughly an additional 0.76 kg of weight loss. Those with two copies of the gene lost around 1.5 kg more than those without. That might sound modest in isolation, but it did represent over 10% of the total weight loss for those with two copies of the GLP1R variant.
A variant in the GIPR gene was linked to increased nausea and vomiting, specifically in people taking tirzepatide. This makes biological sense: tirzepatide is a dual GLP-1/GIP receptor agonist, and the variant is a known partial loss of function mutation. The researchers suggest that when the GIP component can’t function as effectively, it may be less able to buffer the nausea inducing effects of GLP-1 receptor activation. Recent preclinical work supports this theory.
Notably, the GLP1R variant linked to better weight loss was also associated with more nausea and vomiting. And people who were homozygous for risk alleles at both genes faced a roughly 15-fold increased odds of vomiting on tirzepatide. That’s a striking signal for side effect risk.
How much does genetics actually explain?
The study also built a combined prediction model using genetics alongside demographic and clinical factors (age, sex, BMI, drug type, T2D status, and others). The model explained about 25% of the variance in weight loss, with genetics contributing a modest but meaningful slice on top of non-genetic predictors. It was validated against electronic health records from a separate cohort, and people predicted to lose more weight did indeed lose more weight over time.
A few caveats worth noting for your clinical lens. The side effect prediction models performed at 65-68% AUC, which is statistically meaningful but not yet at a level that would change individual patient decisions.
The study also relied heavily on self-reported data from 23andMe participants, who skewed female (82%) and European ancestry (78%). Self-reported weight loss was significantly higher than EHR data showed for the same individuals, so the absolute numbers should be interpreted carefully.
What does this mean for practice right now?
Genetic testing for GLP-1 response isn’t part of standard clinical practice, and we’re not close in terms of actionable, guideline level recommendations. But this study adds weight to the case that personalised approaches to GLP-1 prescribing may be on the horizon. It’s also a useful conversation starter with patients who feel like they’ve “failed” on a GLP-1, when in reality, their response may have been shaped by factors completely outside their control.
You can read the full study in Nature here
References
Su, Q.J., Ashenhurst, J.R., Xu, W., Tran, V., Wu, R.R., Weldon, C.H., Shi, J., Hicks, B., 23andMe Research Team, Abul-Husn, N.S., Aslibekyan, S., Holmes, M.V., Koelsch, B.L. and Auton, A. (2026) ‘Genetic predictors of GLP1 receptor agonist weight loss and side effects’, Nature. doi: 10.1038/s41586-026-10330-z.